A platform for continuously growing biomedical knowledge
The Biomedical Intelligence® Cloud was designed from the ground up to enable learning and growing knowledge from heterogeneous biomedical data and literature.
Biomedical Intelligence Cloud
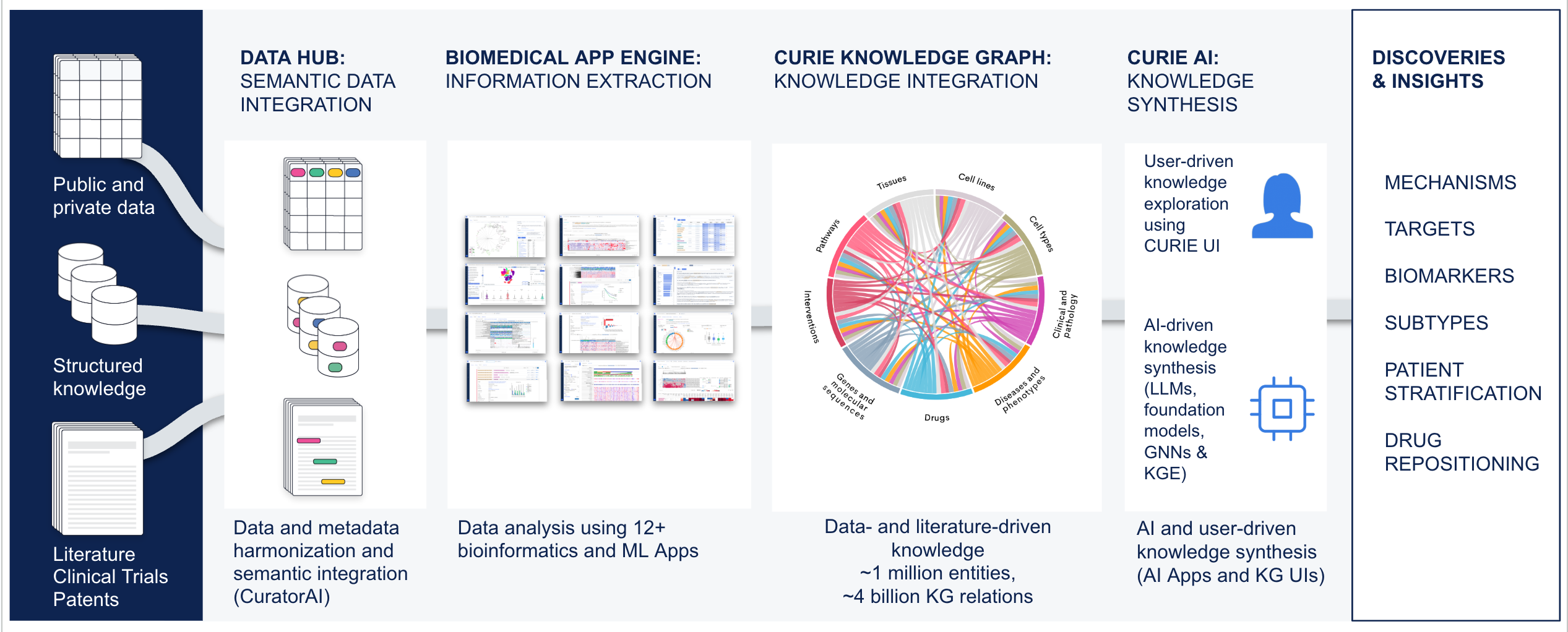
Key Platform Components
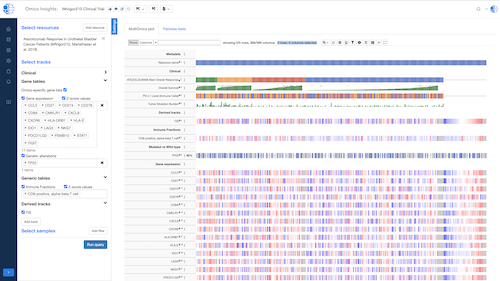
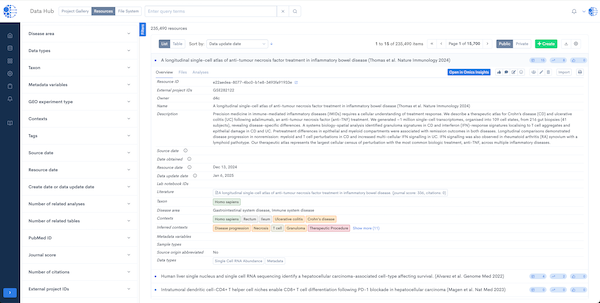
Data Hub – Semantic Data Integration
A semantic data integration and management system that unifies public and private data into a harmonized, searchable repository enriched with detailed metadata.
- Unified Repository: Combines multi-omics, clinical, and literature data in one platform.
- Deep Annotations: Enhances data searchability with semantic metadata.
- Collaboration-Ready: Supports secure data sharing across teams and projects.
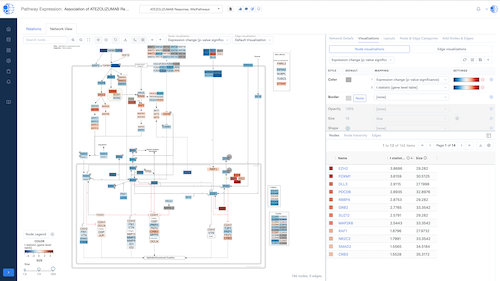
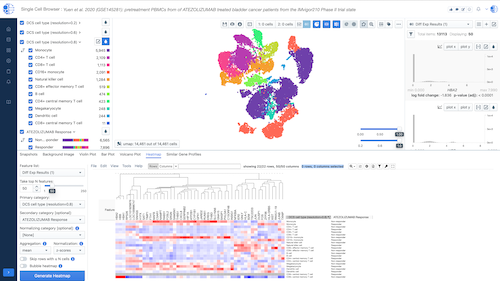
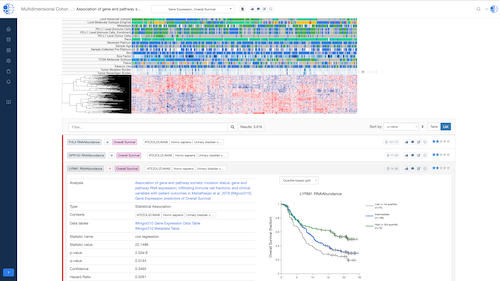
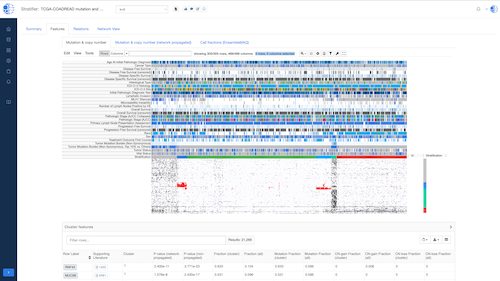
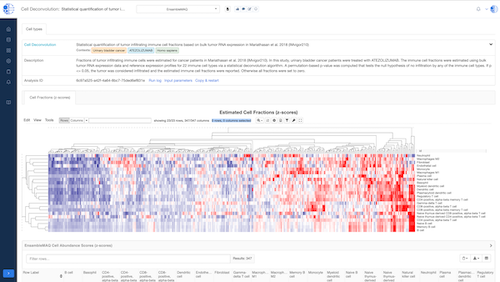
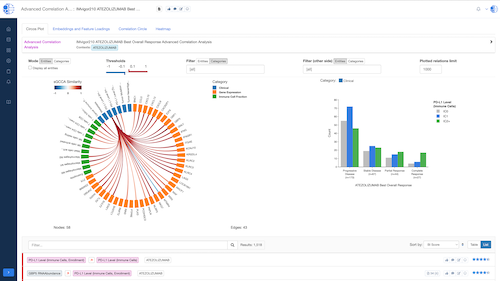
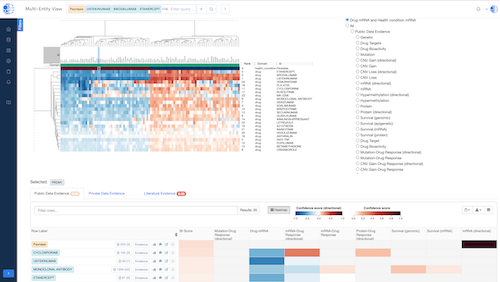
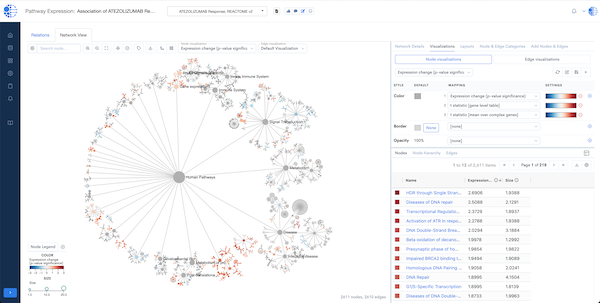
Biomedical App Engine – AI-Powered Analysis Tools
A suite of powerful bioinformatics and machine learning apps designed to extract actionable insights from complex biomedical data.
- Comprehensive Tools: Over 12 apps to analyze your data, including pathway and single-cell analyses, predictive modeling and more.
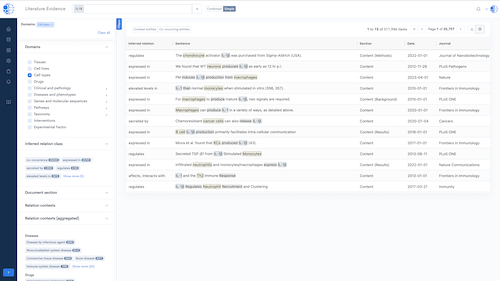
- NLP Engine: Extract insights from literature, clinical trials and patents.
- Knowledge Graph Integration: Leverage the CURIE Knowledge Graph for enriched, context-aware analysis.
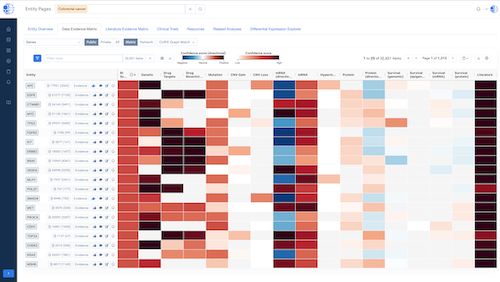
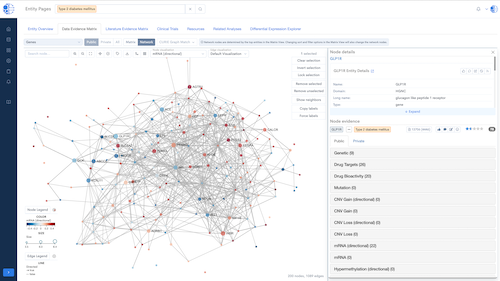
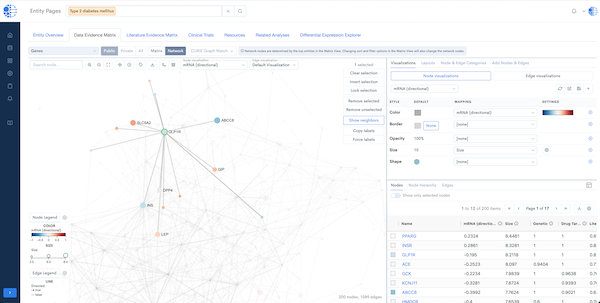
CURIE Knowledge Graph – Knowledge Synthesis at Scale
A continuously updated, data- and literature-driven knowledge graph that synthesizes biomedical knowledge to power discovery.
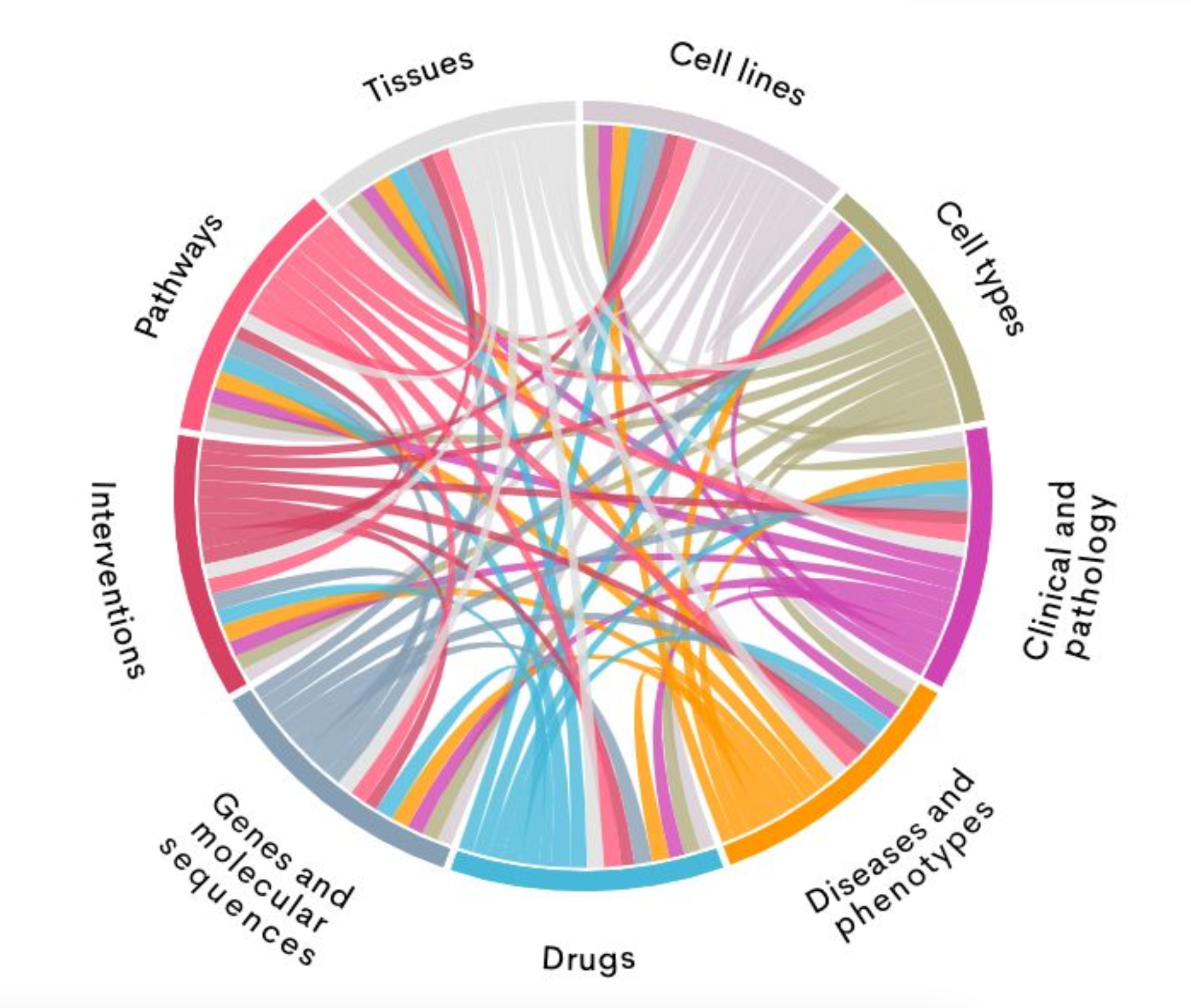
- Comprehensive Knowledge: Over 4 billion data and literature relations connecting genes, diseases, drugs and other biomedical entities.
- Multi-Dimensional Insights: Enables 360° exploration of biomedical entities with evidence from multiple data types.
- Dynamic Updates: Continuously integrates new data and literature.
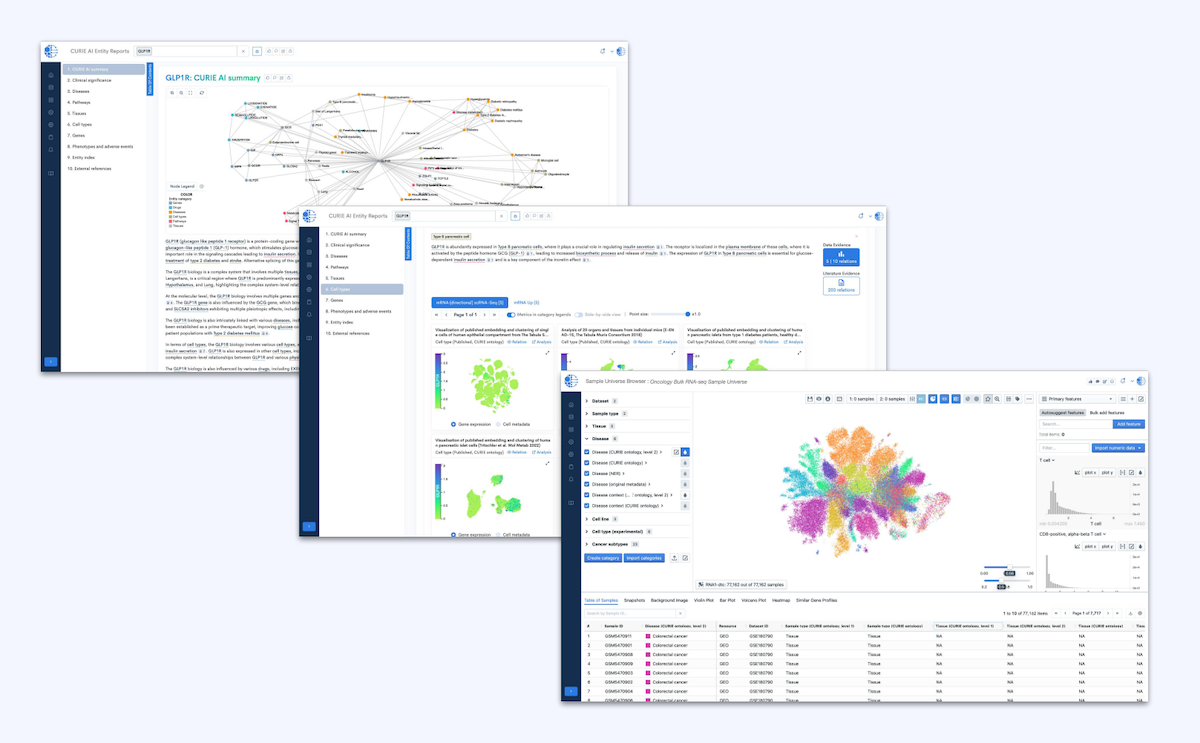
AI Solutions
Purpose-built foundation models and AI-powered tools that drive discoveries and deliver context-rich, explainable insights.
- Predictive Insights: Identify complex biomarkers and response patterns.
- Context-Rich Analysis: Combine structured data and literature for robust findings.
- Interactive Reports: Generate explainable, traceable summaries and scientific narratives.
Biomedical App Engine
Powered by the CURIE Knowledge Graph

Samples 58,555,948

Entities 1,091,114

Analyses 3,277,134

Relations 4,187,567,805
Data4Cure vs Other platforms
Biomedical Intelligence Cloud
Reach Out Today for a Platform DemoOther Platforms
Typical scenario representative of most other platforms in the field
Data4Cure vs Custom development
Biomedical Intelligence Cloud
Reach Out Today for a Platform DemoIn-House Solution
Using custom pipelines and applications or combining multiple third-party solutions